Quantum State Tomography¶
Quantum tomography is an experimental procedure to reconstruct a description of part of a quantum system from the measurement outcomes of a specific set of experiments. In particular, quantum state tomography reconstructs the density matrix of a quantum state by preparing the state many times and measuring them in a tomographically complete basis of measurement operators.
Note
This tutorial requires the qiskit-aer and qiskit-ibm-runtime
packages to run simulations. You can install them with python -m pip
install qiskit-aer qiskit-ibm-runtime.
We first initialize a simulator to run the experiments on.
from qiskit_aer import AerSimulator
from qiskit_ibm_runtime.fake_provider import FakePerth
backend = AerSimulator.from_backend(FakePerth())
To run a state tomography experiment, we initialize the experiment with a circuit to
prepare the state to be measured. We can also pass in an
Operator or a Statevector
to describe the preparation circuit.
import qiskit
from qiskit_experiments.framework import ParallelExperiment
from qiskit_experiments.library import StateTomography
# GHZ State preparation circuit
nq = 2
qc_ghz = qiskit.QuantumCircuit(nq)
qc_ghz.h(0)
qc_ghz.s(0)
for i in range(1, nq):
qc_ghz.cx(0, i)
# QST Experiment
qstexp1 = StateTomography(qc_ghz)
qstdata1 = qstexp1.run(backend, seed_simulation=100).block_for_results()
# Print results
display(qstdata1.analysis_results(dataframe=True))
| name | experiment | components | value | quality | backend | run_time | trace | eigvals | raw_eigvals | rescaled_psd | fitter_metadata | conditional_probability | positive | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 9a4002b0 | state | StateTomography | [Q0, Q1] | DensityMatrix([[ 0.4749349 +0.j , -0.00... | unknown | aer_simulator_from(fake_perth) | None | 1.0 | [0.9137554587098121, 0.04180368599957844, 0.02... | [0.9137554587098121, 0.04180368599957844, 0.02... | False | {'fitter': 'linear_inversion', 'fitter_time': ... | 1.0 | True |
| 0a328501 | state_fidelity | StateTomography | [Q0, Q1] | 0.913086 | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
| 02b2147b | positive | StateTomography | [Q0, Q1] | True | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
Tomography Results¶
The main result for tomography is the fitted state, which is stored as a
DensityMatrix object:
state_result = qstdata1.analysis_results("state", dataframe=True).iloc[0]
print(state_result.value)
DensityMatrix([[ 0.4749349 +0.j , -0.00813802+0.01204427j,
0.00764974-0.00227865j, 0.00048828-0.44042969j],
[-0.00813802-0.01204427j, 0.03222656+0.j ,
-0.00048828+0.00390625j, -0.00406901+0.00455729j],
[ 0.00764974+0.00227865j, -0.00048828-0.00390625j,
0.02246094+0.j , 0.01432292-0.01432292j],
[ 0.00048828+0.44042969j, -0.00406901-0.00455729j,
0.01432292+0.01432292j, 0.4703776 +0.j ]],
dims=(2, 2))
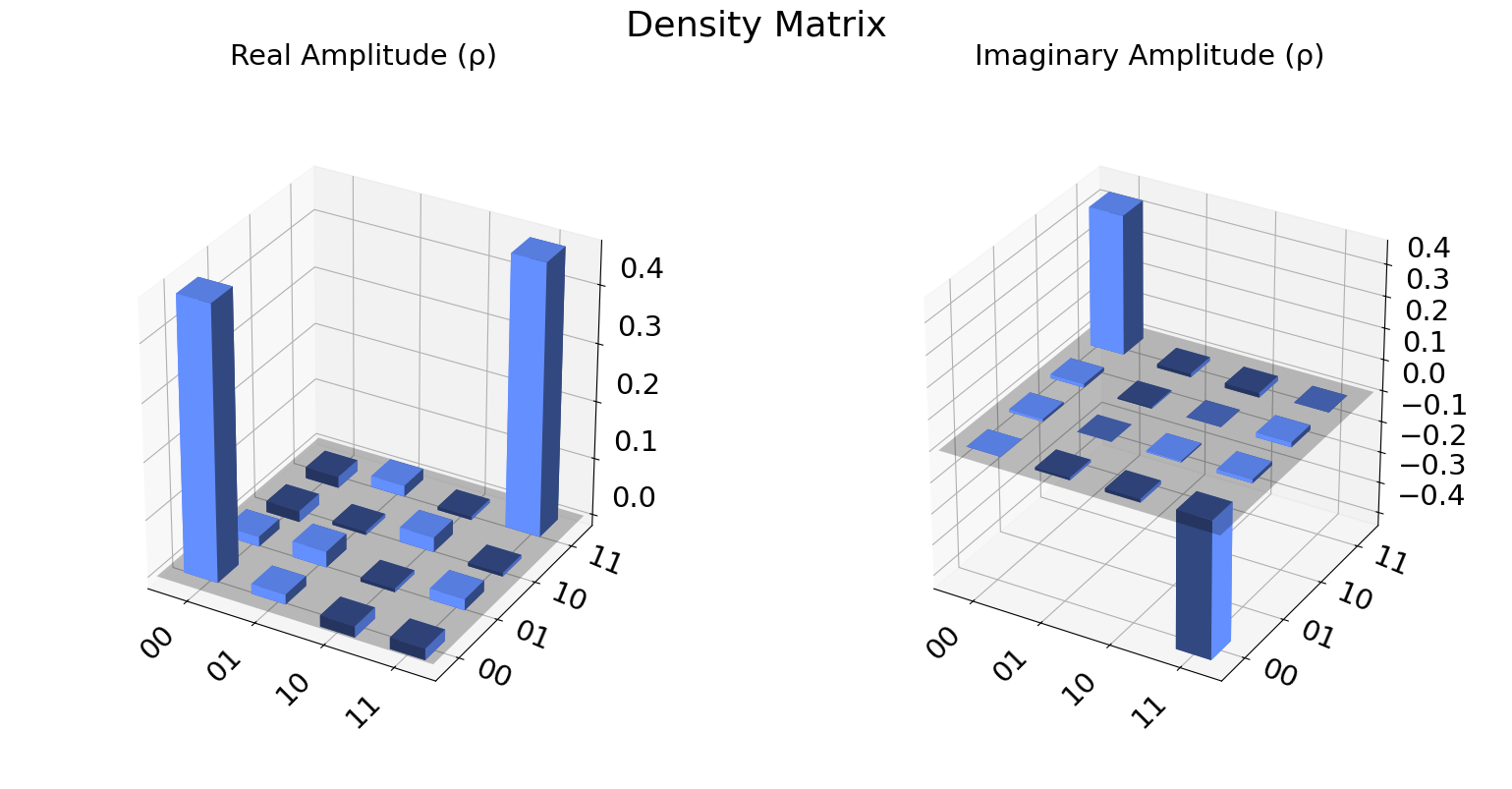
We can also visualize the density matrix:
from qiskit.visualization import plot_state_city
state = qstdata1.analysis_results("state", dataframe=True).iloc[0].value
plot_state_city(state, title='Density Matrix')
The state fidelity of the fitted state with the ideal state prepared by
the input circuit is stored in the "state_fidelity" result field.
Note that if the input circuit contained any measurements the ideal
state cannot be automatically generated and this field will be set to
None.
fid_result = qstdata1.analysis_results("state_fidelity", dataframe=True).iloc[0]
print("State Fidelity = {:.5f}".format(fid_result.value))
State Fidelity = 0.91309
Additional state metadata¶
Additional data is stored in the tomography under additional fields. This includes
eigvals: the eigenvalues of the fitted statetrace: the trace of the fitted statepositive: Whether the eigenvalues are all non-negative
If trace rescaling was performed this dictionary will also contain a raw_trace field
containing the trace before rescaling. Futhermore, if the state was rescaled to be
positive or trace 1 an additional field raw_eigvals will contain the state
eigenvalues before rescaling was performed.
for col in ["eigvals", "trace", "positive"]:
print(f"{col}: {state_result[col]}")
eigvals: [0.91375546 0.04180369 0.02965482 0.01478603]
trace: 1.0000000000000016
positive: True
To see the effect of rescaling, we can perform a “bad” fit with very low counts:
# QST Experiment
bad_data = qstexp1.run(backend, shots=10, seed_simulation=100).block_for_results()
bad_state_result = bad_data.analysis_results("state", dataframe=True).iloc[0]
# Print result
for key, val in bad_state_result.items():
print(f"{key}: {val}")
name: state
experiment: StateTomography
components: [<Qubit(Q0)>, <Qubit(Q1)>]
value: DensityMatrix([[ 0.33450396+0.j , 0.05938614-0.04566114j,
-0.11306592+0.05611775j, 0.05798364-0.39626276j],
[ 0.05938614+0.04566114j, 0.0436272 +0.j ,
-0.06379144+0.00282676j, 0.09825567-0.06775048j],
[-0.11306592-0.05611775j, -0.06379144-0.00282676j,
0.09861805+0.j , -0.13320385+0.12088397j],
[ 0.05798364+0.39626276j, 0.09825567+0.06775048j,
-0.13320385-0.12088397j, 0.52325079+0.j ]],
dims=(2, 2))
quality: unknown
backend: aer_simulator_from(fake_perth)
run_time: None
trace: 1.0000000000000007
eigvals: [0.92575281 0.07424719 0. 0. ]
raw_eigvals: [ 1.05200742 0.20050181 -0.06354245 -0.18896677]
rescaled_psd: True
fitter_metadata: {'fitter': 'linear_inversion', 'fitter_time': 0.0043561458587646484}
conditional_probability: 1.0
positive: True
Tomography Fitters¶
The default fitters is linear_inversion, which reconstructs the
state using dual basis of the tomography basis. This will typically
result in a non-positive reconstructed state. This state is rescaled to
be positive-semidefinite (PSD) by computing its eigen-decomposition and
rescaling its eigenvalues using the approach from Ref. [1].
There are several other fitters are included (See API documentation for
details). For example, if cvxpy is installed we can use the
cvxpy_gaussian_lstsq() fitter, which allows constraining the fit to be
PSD without requiring rescaling.
try:
import cvxpy
# Set analysis option for cvxpy fitter
qstexp1.analysis.set_options(fitter='cvxpy_gaussian_lstsq')
# Re-run experiment
qstdata2 = qstexp1.run(backend, seed_simulation=100).block_for_results()
state_result2 = qstdata2.analysis_results("state", dataframe=True).iloc[0]
for key, val in state_result2.items():
print(f"{key}: {val}")
except ModuleNotFoundError:
print("CVXPY is not installed")
name: state
experiment: StateTomography
components: [<Qubit(Q0)>, <Qubit(Q1)>]
value: DensityMatrix([[ 0.47360887+0.j , 0.01651754-0.01645398j,
0.00528622-0.00449668j, -0.01767143-0.43915202j],
[ 0.01651754+0.01645398j, 0.03051735+0.j ,
0.00677056+0.00852105j, -0.00747277+0.00333155j],
[ 0.00528622+0.00449668j, 0.00677056-0.00852105j,
0.03197703+0.j , -0.01182688+0.0033077j ],
[-0.01767143+0.43915202j, -0.00747277-0.00333155j,
-0.01182688-0.0033077j , 0.46389675+0.j ]],
dims=(2, 2))
quality: unknown
backend: aer_simulator_from(fake_perth)
run_time: None
trace: 0.9999999972176197
eigvals: [0.908474 0.05950643 0.02763186 0.0043877 ]
raw_eigvals: [0.90847401 0.05950643 0.02763186 0.0043877 ]
rescaled_psd: False
fitter_metadata: {'fitter': 'cvxpy_gaussian_lstsq', 'cvxpy_solver': 'SCS', 'cvxpy_status': ['optimal'], 'psd_constraint': True, 'trace_preserving': True, 'fitter_time': 0.037584781646728516}
conditional_probability: 1.0
positive: True
Parallel Tomography Experiment¶
We can also use the ParallelExperiment class to
run subsystem tomography on multiple qubits in parallel.
For example if we want to perform 1-qubit QST on several qubits at once:
from math import pi
num_qubits = 5
gates = [qiskit.circuit.library.RXGate(i * pi / (num_qubits - 1))
for i in range(num_qubits)]
subexps = [
StateTomography(gate, physical_qubits=(i,))
for i, gate in enumerate(gates)
]
parexp = ParallelExperiment(subexps)
pardata = parexp.run(backend, seed_simulation=100).block_for_results()
display(pardata.analysis_results(dataframe=True))
| name | experiment | components | value | quality | backend | run_time | trace | eigvals | raw_eigvals | rescaled_psd | fitter_metadata | conditional_probability | positive | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 503e0659 | state | StateTomography | [Q0] | DensityMatrix([[0.96777344+0.j , 0.0175... | unknown | aer_simulator_from(fake_perth) | None | 1.0 | [0.9682268402188912, 0.03177315978110963] | [0.9682268402188912, 0.03177315978110963] | False | {'fitter': 'linear_inversion', 'fitter_time': ... | 1.0 | True |
| 0bf4ab4b | state_fidelity | StateTomography | [Q0] | 0.967773 | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
| da1fe881 | positive | StateTomography | [Q0] | True | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
| 471ee0e1 | state | StateTomography | [Q1] | DensityMatrix([[0.82910156+0.j , 0.0097... | unknown | aer_simulator_from(fake_perth) | None | 1.0 | [0.9607150300654657, 0.03928496993453533] | [0.9607150300654657, 0.03928496993453533] | False | {'fitter': 'linear_inversion', 'fitter_time': ... | 1.0 | True |
| 98e3e436 | state_fidelity | StateTomography | [Q1] | 0.960586 | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
| 22cef7f8 | positive | StateTomography | [Q1] | True | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
| 04411412 | state | StateTomography | [Q2] | DensityMatrix([[0.5234375+0.j , 0.00781... | unknown | aer_simulator_from(fake_perth) | None | 1.0 | [0.9684253840187003, 0.03157461598130054] | [0.9684253840187003, 0.03157461598130054] | False | {'fitter': 'linear_inversion', 'fitter_time': ... | 1.0 | True |
| 96efb51e | state_fidelity | StateTomography | [Q2] | 0.967773 | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
| 15272b05 | positive | StateTomography | [Q2] | True | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
| ce35f27a | state | StateTomography | [Q3] | DensityMatrix([[ 0.19238281+0.j , -0.013671... | unknown | aer_simulator_from(fake_perth) | None | 1.0 | [0.9387157442028106, 0.061284255797190226] | [0.9387157442028106, 0.061284255797190226] | False | {'fitter': 'linear_inversion', 'fitter_time': ... | 1.0 | True |
| 5de0d93d | state_fidelity | StateTomography | [Q3] | 0.938489 | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
| 1ad60c4c | positive | StateTomography | [Q3] | True | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
| e1ba49f4 | state | StateTomography | [Q4] | DensityMatrix([[0.03320313+0.j , 0. ... | unknown | aer_simulator_from(fake_perth) | None | 1.0 | [0.9668224120237543, 0.033177587976246654] | [0.9668224120237543, 0.033177587976246654] | False | {'fitter': 'linear_inversion', 'fitter_time': ... | 1.0 | True |
| c84e5c71 | state_fidelity | StateTomography | [Q4] | 0.966797 | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
| c9709ff8 | positive | StateTomography | [Q4] | True | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
View experiment analysis results for one component:
results = pardata.analysis_results(dataframe=True)
display(results[results.components.apply(lambda x: x == ["Q0"])])
| name | experiment | components | value | quality | backend | run_time | trace | eigvals | raw_eigvals | rescaled_psd | fitter_metadata | conditional_probability | positive | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 503e0659 | state | StateTomography | [Q0] | DensityMatrix([[0.96777344+0.j , 0.0175... | unknown | aer_simulator_from(fake_perth) | None | 1.0 | [0.9682268402188912, 0.03177315978110963] | [0.9682268402188912, 0.03177315978110963] | False | {'fitter': 'linear_inversion', 'fitter_time': ... | 1.0 | True |
| 0bf4ab4b | state_fidelity | StateTomography | [Q0] | 0.967773 | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
| da1fe881 | positive | StateTomography | [Q0] | True | unknown | aer_simulator_from(fake_perth) | None | None | None | None | None | None | None | None |
References¶
See also¶
API documentation:
StateTomography